Protein phosphorylation is a key regulatory mechanism in most cellular processes. In eukaryotes, it is predominantly the amino acids serine (S), threonine (T) and tyrosine (Y) that are modified by covalent attachment of a phosphate group. This often takes place on multiple sites on a given substrate. It is a highly dynamic process, with phosphate addition catalysed by enzymes called (protein) kinases and removal catalysed by enzymes called (phosphoprotein) phosphatases. The phosphate donor is usually ATP, the cell's energy currency, which is continually re-synthesised by core metabolic processes.
A substrate molecule with n sites could have N = 2n global patterns of phosphorylation, or "phospho-forms". Different phospho-forms may have different effects, so that the behaviour of a multisite phosphorylation system depends on its "phospho-form distribution". The combinatorial explosion of phospho-forms makes it challenging to analyse this distribution both experimentally and mathematically
Suppose that a n-site substrate is mixed with kinase and phosphatase, with ATP not limiting, and allowed to reach a quasi-steady state, in which the concentrations of the N phospho-forms are measured and plotted as a point in N-dimensional space. Suppose this is done repeatedly, for different total amounts of substrate and enzymes and different starting conditions of substrate. What shape is the resulting set of points?
Surprisingly, theory predicts that it is a curve, or 1-dimensional algebraic variety, irrespective of n [♦]. Indeed, the curve can be rationally parameterised by a single auxiliary variable, which makes it very special. The rational parameterisation implies that the steady states of the entire system can be calculated by solving just 2 algebraic equations, instead of having to numerically integrate an exponential number of differential equations. We can mathematically analyse such systems without having to know in advance how many sites they have or their parameter values. We might hope, in this way, to see principles of biological regulation emerge from the molecular complexity in which they are implemented.
One such result comes from observing that as n increases, while the dimension of the phospho-form variety does not change, its geometry does: it can "wiggle" more. This means that the amount of information that can, in principle, be encoded in the system, increases with n [♦]. Perhaps this is one reason for the large numbers of sites on some key proteins, particularly in metazoan organisms.
Phosphorylation is only one of several protein post-translational modifications, which include methylation, acteylation, ubiquitination, GlcNAcylation, …. These work together to regulate cellular processes. The rational parmeterisation result holds for multiple enzymes, carrying out multiple types of modification on multiple substrates. The steady state "mod-forms" are parameterised by p variables, where p is the number of enzymes, irrespective of the number of substrates or sites. Instead of a curve, we get a variety of dimension no more than p [♦].
previous next back to research
 |
|
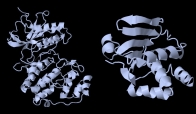
(Left) Ribbon diagram of the crystal structure of the S/T kinase Erk2, showing the characteristic two lobes with the ATP binding pocket in between. PDB 1erk. (Right) The dual-specificity phosphatase MKP3, which is able to remove both S/T and Y phosphates. PDB 1mkp |
 |
|
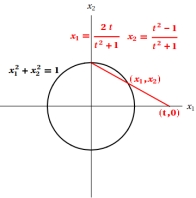
A circle can be defined as an algebraic variety either implicitly, as the solutions of a polynomial equation, or explicitly, through a parameterisation by rational functions, as shown. In fact, all conic sections can be rationally parameterised in this way. For higher degree curves, the question of whether they are rational or not is much more subtle and working out the answer was one of the triumphs of 19th century mathematics. Rationality depends on a non-negative integer, called the genus of the curve. Glossing over some important issues, a curve is rational if, and only if, its genus is zero. If d is the degree of a curve, then genus = (d-1)(d-2)/2 - ∑c(p) where p runs over the singular points of the curve and c(p) is a non-negative integer that measures the extent of singularity. The more singularities, the lower the genus. That is why all the curves on the previous page are rational except for the non-singular cubic, whose genus is 1. |