We are developing mass spectometry techniques to measure phospho-form distributions. This will enable us to undertake the in-vitro thought experiment on the previous page, with real substrates and enzymes. The rational parameterisation theorem predicts an upper bound for the dimension of the resulting set of points. Can we deduce anything further?
Consider a kinase, phosphatase, substrate system with 2 sites, in which both enzymes act distributively, so that at most one modification (either phosphorylation or dephosphorylation) is done at a time. We can show that the steady state concentrations of the four phospho-forms satisfies the quadratic formula
a[S01]2 + b[S01][S10] + c[S10]2 = d[S00][S11]
where a, b, c, d depend only on the parameters [♦]. This formula holds for any steady state, irrespective of the total amounts of substrate and enzymes and the starting conditions. We call it an invariant. Dividing by the term on the right and treating the expression as a linear form in three new variables, we see that the phospho-form variety, in these new variables, lies on a plane. This condition can be checked without having to know anything at all about the parameter values (which is particularly important because they are almost never known!).
If one or the other of the enzymes is processive, so that more than one modification occurs at a time, then the planarity condition no longer holds. However, the phospho-form variety has a different geometry in each of the four possible cases. We can therefore infer details of the underlying biochemical network by simply inspecting the geometry of the steady-state phospho-form variety.
Invariants are algebraic consequences of the polynomials in the system's ODEs, which are useful because they do not involve certain variables, particularly the free enzyme concentrations, which are extremely difficult to measure. Algebraic geometry provides the Gröbner basis method for systematically eliminating variables from polynomial equations [♦]. This algorithm is implemented in Mathematica and is one way to find invariants. For an elementary overview of algebraic geometry and Gröbner bases, see [♦].
Guy Shinar and Uri Alon have introduced "absolute concentration robustness" [♦]. A system exhibits this if there is an invariant of the form [X] = a, so that the concentration of X has the same value at steady state, no matter how the system is initiated. This robustness has been experimentally observed in certain bacterial metabolic systems [♦,♦]. An invariant can encode a more complex form of robustness, in which several variables may participate.
 |
|
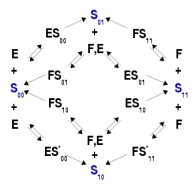
A biochemical network for a 2-site substrate, S, acted on by a kinase, E, and a phosphatase, F. The 4 phospho-forms are shown in blue, with the subscript bitstring showing which sites are phosphorylated. ATP is assumed to be maintained constant by processes which are not explicitly modelled and its affect absorbed into the rate constants. The enzymes are assumed to act distributively; a processive enzyme would need an additional reaction S00 <------> S11 in which two modifications happen at once [Fig 3b of ♦]. |
 |
|
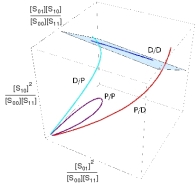
When the enzymes are distributive (D/D), the steady state variety (blue) lies on a plane (cyan) in the new coordinates coming from the invariant formula. We already know that this variety is a curve because of the rational parameterisation theorem on the previous page. If one or more of the enzymes are processive, the curve converges to the origin for small (D/P), large (P/D) or both (P/P) values of the rational parameter [♦]. |