Mathematical models are increasingly used to understand the behaviour of molecular networks. A very common type of model consists of a dynamical system, specified by ordinary differential equations (ODEs)
dx/dt = f(x;a)
Here, x is a vector of state variables describing the concentrations of each of the molecular components in the network and f describes the net rate of production of each variable, which may be a function of the other variables. These rates usually depend on various "rate constants", or "parameters", which are collected together in the vector a. The complexity of molecular networks is such that the sizes of both x and a can be very large.
The most systematic way of constructing such dynamical systems is to generate them from the underlying biochemical reactions using the Principle of Mass Action. Suppose that we have a hypothetical (bio)chemical reaction
2A + 3B ---> C
(Very few actual reactions have such stoichiometry but this illustrates the idea). If this reaction is elementary, so that no intermediates are formed, then the rate of production of C is proportional to the product of the concentrations of the substrates to the power of their respective stoichiometries. In other words,
d[C]/dt ---> k[A]2[B]3,
where k is the constant of proportionality, or "rate constant". This principle seems to work well in the test tube, although a modern chemist might replace concentration with chemical activity. It remains unclear whether the principle works equally well inside cells because the kinds of data that we can currently gather are not good enough to tell. However, models constructed on the basis of mass action have been biologically informative and the principle has the great advantage of being a systematic method that can be applied to any set of elementary reactions.
previous next back to research
 |
|
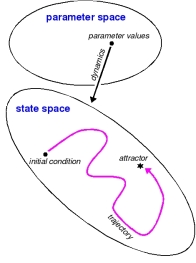
Schematic of a dynamical system. A choice of parameter values defines a dynamics on the state space. Given an initial condition, the dynamics determines a trajectory of the system. This may lead to a steady state, as shown, or to a more complex attractor. [♦] |
 |
|
Cato Guldberg and Peter Waage, who first stated the Principle of Mass Action in 1864. Guldberg was a mathematician and Waage a chemist, whose second wife was Guldberg's sister. |